Cluster treeview11/3/2022  
Select clusterMaker and click the Install button.įigure 2. Select Analysis under Available for Install. Is available in the Analysis group of plugins. You must be running Cytoscape 2.8.2 or newer. To download clusterMaker using the plugin manager, Create New Network with Nested Networks from AttributeĬlusterMaker is available through the Cytoscape plugin manager orīy downloading the source directly from the Cytoscape svn repositoryĬytoscape Subversion Server information, or browse the csplugins/ucsf/scooter/clusterMaker.SCPS (Spectral Clustering of Protein Sequences).Gene expression analysis in a network context.įinding complexes in proteomic and genetic interaction data.įunctional annotation by clustering protein similarity networks. In the BMC Bioinformatics publication: " clusterMaker: A Multi-algorithm Clustering Plugin for Cytoscape", currently submitted. The other three tutorials describe the steps necessary to reproduce the scenarios described The first tutorial:Ĭluster Maker covers some of the basic featuresĪnd uses of clusterMaker. Open Tutorials web site under the "Cytoscape Tutorials" section. In addition to this documentation, there are four tutorials available on the "meta nodes" to allow interactive exploration of the putative familyĪssociations within the Cytoscape network, and results may alsoīe shown as a separate network containing only the intra-cluster edges, or with inter-cluster edges added back.ĬlusterMaker requires version 2.8.2 or newer of CytoscapeĪnd is available from the Cytoscape plugin manager under Hierarchical, k-medoid, AutoSOME, and k-means clusters may beĭisplayed as hierarchical groups of nodes or as heat maps.Īll of the network partitioning cluster algorithms create collapsible MCODE, community clustering (GLAY), SCPS, and AutoSOME for partitioning networks based on similarity K-means for clustering expression or genetic data and MCL, transitivity clustering, affinity propagation, Current clustering algorithms include hierarchical, k-medoid, AutoSOME, and UCSF clusterMaker is a Cytoscape plugin that unifiesĭifferent clustering techniques and displays into a single In this screenshot, theĮxpression data in the sampleData file galFiltered.cys has been clustered using the hierarchical methodĪnd displayed as a heatmap with associated dendrogram. CLUSTER TREEVIEW FULLSee the source of Cluster.pm for a full copyright statement.Figure 1. This module is a Perl wrapper for the C clustering library for cDNA microarray data, Copyright (C) 2002 Michiel Jan Laurens de Hoon. A copyright statement is contained in the source code itself. CLUSTER TREEVIEW SOFTWAREThanks to Michael Eisen, for creating the software packages Cluster and TreeView. See the scripts in the examples subdirectory of the package. The C Clustering Library is distributed along with Cluster 3.0, an enhanced version of the famous Cluster program originally written by Michael Eisen while at Stanford University. This library was developed at the Human Genome Center of the University of Tokyo. This module is an interface to the C Clustering Library, a general purpose library implementing functions for hierarchical clustering (pairwise simple, complete, average, and centroid linkage), along with k-means and k-medians clustering, and 2D self-organizing maps. 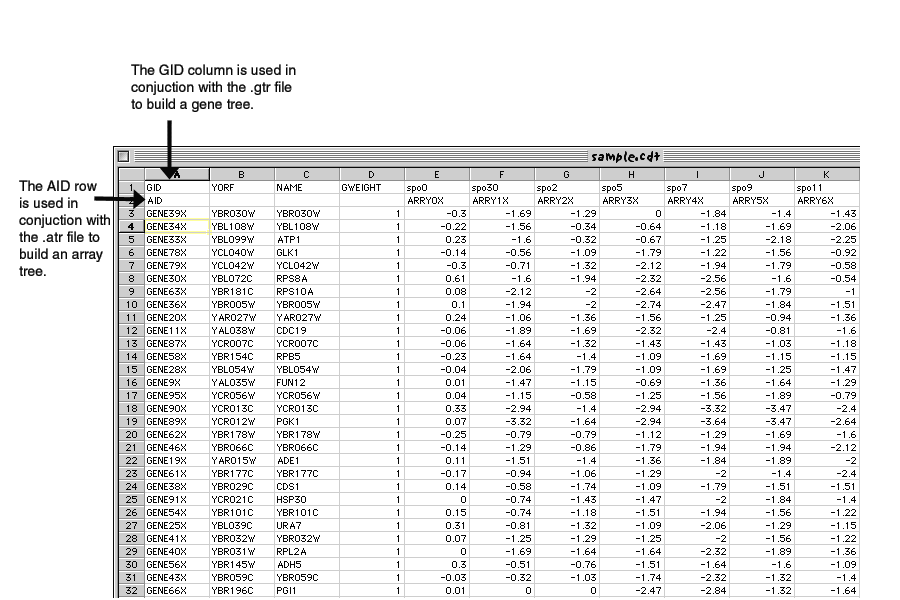
Algorithm::Cluster - Perl interface to the C Clustering Library. 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
